Home > Teams > Metabolic and Cardiovascular Diseases > HCEMM-USZ Translational Pancreatology Research Group
Group Leader: Balazs Németh M.D., Ph.D.
His junior group seeks possible treatment options of chronic pancreatitis, that is a progressive, relapsing inflammatory disorder of the pancreas. This disease often develops after multiple episodes of acute pancreatitis in the background of chronic alcoholism or genetic predisposition. Since specific therapy is still lacking, animal models that faithfully mimic human chronic pancreatitis both phenotypically and mechanistically are essential for the development of new therapeutic and preventive measures. In our HCEMM project we use a novel preclinical mouse model of pancreatitis driven by trypsinogen misfolding. Trypsinogen, the most abundant digestive enzyme produced by the pancreas plays a key role in developing pancreatitis. However, early trypsinogen activation inside the pancreas is the major biochemical effect in this disease, there are some other known pathways that may induce chronic pancreatitis. Protein misfolding and unfolded protein response caused by mutations in pancreas specific enzymes (CPA1, CTRC, PRSS1 and CEL) eventually lead to the development of chronic pancreatitis independently form trypsinogen activation. Experimental modelling of this mechanism may help us to better understand the disease development and gives the possibility to test several compounds that may diminish or slow down the progression of the disease. Collaboration with the Hungarian Pancreatic Study Group gives us the chance to continuously participate in clinical studies of pancreatitis. As a result of this collaboration, a significant part of our scientific activity is the biochemical characterization and functional analysis of novel variants found in pancreatitis associated genes in patients with pancreatitis.
Group Leader: László Csanády MD, PhD, DSc
Our group studies structure and function of two medically relevant ion channels, the Cystic Fibrosis Transmembrane Conductance Regulator (CFTR) anion channel and the Ca2+ permeable cation channel Transient Receptor Potential Melastatin 2 (TRPM2). CFTR and TRPM2 are known as “channel-enzymes”, proteins in which a single polypeptide chain displays both ion channel and enzymatic properties. Because impaired or excessive activity of either channel causes disease, both are promising therapeutic targets: understanding their molecular mechanisms would be an important conceptual, development of specific activators/inhibitors an outstanding practical, achievement.
CFTR hyperactivity leads to diarrhea (e.g., cholera), whereas its mutations cause cystic fibrosis (CF), the most common lethal inherited disease among whites. A mutation found in 90% of CF patients impairs both CFTR surface expression and channel gating: effective therapy will thus require a combination of a “corrector” drug to enhance surface expression, and of a “potentiator” to stimulate gating. Our goals are to deepen our understanding of CFTR structure, physiological regulation, and the molecular motions that underlie channel gating. We further wish to dissect the molecular pathologies of CFTR mutations that cause CF. Finally, we hope to exploit this knowledge to promote mechanism-guided potentiator design.
TRPM2 activity is needed for the immune response, insulin secretion, and the maintanance of body temperature, but its excessive activation after ischemia causes cell death: thus, both TRPM2 inhibition (stroke, heart attack, Alzheimer’s, inflammation, hyperinsulinism) and stimulation (diabetes, amyotrophic lateral sclerosis, Parkinsonism, bipolar disorder, excessive fever) might become clinically useful. Our aim is to better understand TRPM2 structure, function, and evolutionary adaptation in vertebrates, as well as a detailed pharmacological profiling of the human channel.
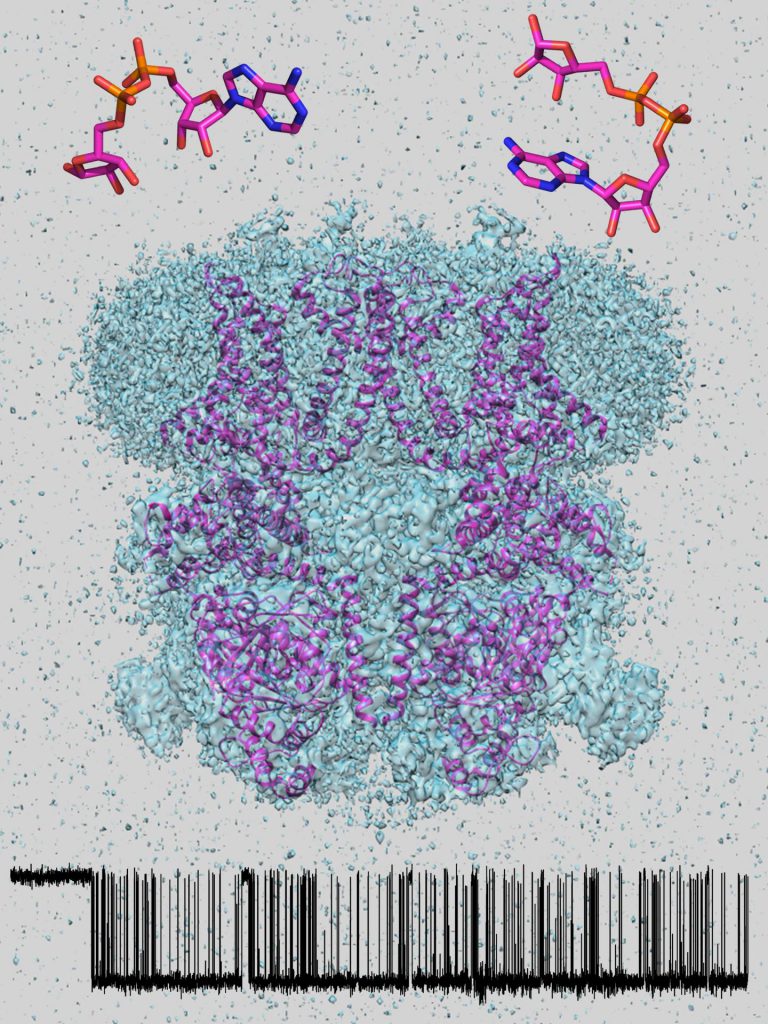
Our methodology includes high-resolution electrophysiological (patch-clamp) recordings of channel currents from native cells and heterologous expression systems, in vitro enzymatic assays on heterologously expressed channel proteins purified to homogeneity, electron cryomicroscopy, and in silico structure modeling.